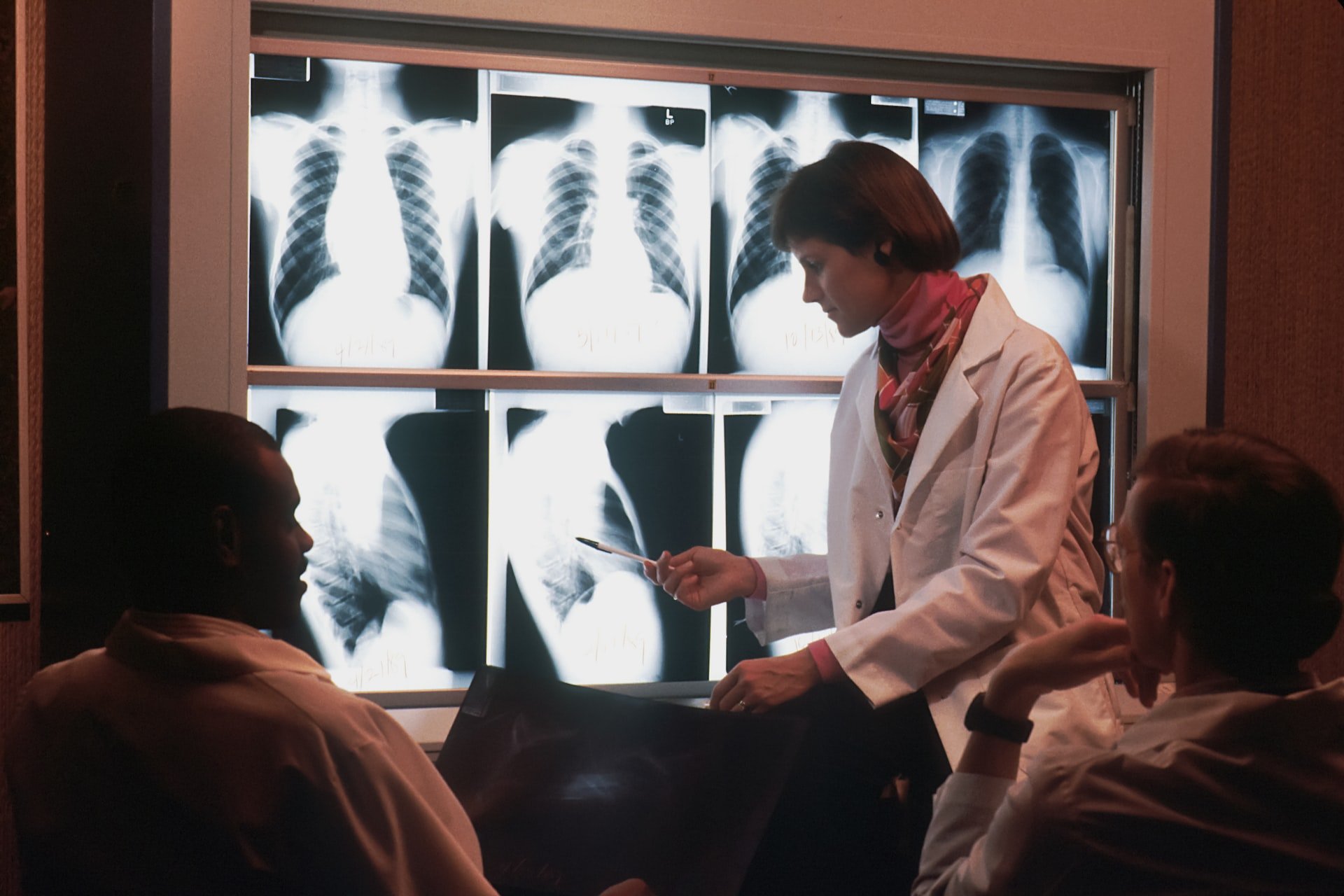
AI systems in healthcare often rely on human-labeled data sets. A new method overcomes this limitation with self-supervised learning for X-ray images.
AI systems for medical diagnosis of chest X-rays help radiologists detect lung disease or cancer. To train those AI models, researchers resort to supervised learning algorithms. This training method has proven itself in other areas of image analysis and is considered reliable.
But supervised learning requires data labeled by humans. In medicine, this is a major problem: interpreting medical images is time-consuming and requires expert radiologists. Data is therefore a bottleneck for improving and extending AI diagnostics to other diseases. Some researchers are therefore already experimenting with AI models that are pre-trained with unlabeled data and then fine-tuned with the rare data labeled by experts.
Self-supervised trained models are on the way to SOTA
Outside medical diagnostics, meanwhile, self-supervised trained models have become prevalent for image tasks. Early examples such as Meta's SEER or DINO and Google's Vision Transformer showed the potential of combining self-supervised training methods with huge, unlabeled datasets.
Breakthroughs such as OpenAI's CLIP highlighted the central role models trained multimodal with images and associated text descriptions could play in image analysis and generation. In April, Google demonstrated a multimodal model using the LiT method that achieved the quality of supervised trained models in the ImageNet benchmark.
Researchers at Harvard Medical School and Stanford University have now made a major advance: They are releasing CheXzero, a self-supervised trained AI model that autonomously learns to identify diseases from images and text data.
CheXzero learns from X-rays and clinical records
The researchers trained CheXzero with more than 377,000 chest X-rays and more than 227,000 associated clinical notes. The team then tested the model with two separate datasets from two different institutions, one outside the U.S., to ensure robustness with different terminology.
In the tests, ChexZero successfully identified pathologies that were not explicitly mentioned in the notes made by human experts. The model outperformed other self-supervised AI tools and approached the accuracy of human radiologists.
"We're living the early days of the next-generation medical AI models that are able to perform flexible tasks by directly learning from text," said study lead investigator Pranav Rajpurkar, assistant professor of biomedical informatics in the Blavatnik Institute at HMS. "Up until now, most AI models have relied on manual annotation of huge amounts of data—to the tune of 100,000 images—to achieve a high performance. Our method needs no such disease-specific annotations."
CheXzero removes data bottleneck for medical AI systems
CheXzero learns how concepts in the unstructured text correspond to visual patterns in images, Rajpurkar said. The method is, therefore, suitable for other imaging modalities, such as CT scans, MRIs, and echocardiography.
"CheXzero shows that accuracy of complex medical image interpretation no longer needs to remain at the mercy of large, labeled datasets," said study co-first author Ekin Tiu, an undergraduate student at Stanford and a visiting researcher at HMS. "We use chest X-rays as a driving example, but in reality CheXzero's capability is generalizable to a vast array of medical settings where unstructured data is the norm, and precisely embodies the promise of bypassing the large-scale labeling bottleneck that has plagued the field of medical machine learning"
The code and model will be made publicly available on GitHub and will be accompanied by a scientific publication in Nature.